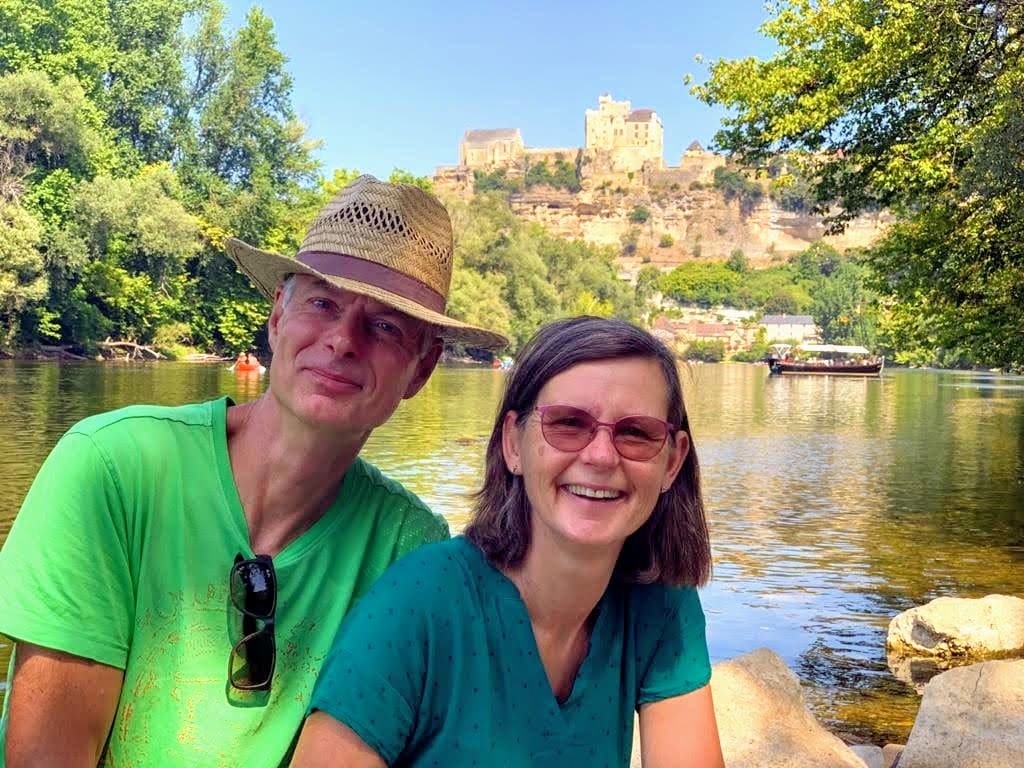

Verslavingspreventie: het spotten van risicofactoren
Wanneer we denken aan verslaving, komt vaak het beeld van een donker pad voor ogen


Wanneer we denken aan verslaving, komt vaak het beeld van een donker pad voor ogen
Als je denkt aan het verbeteren van je fysieke fitheid, denk je waarschijnlijk meteen aan

Als je droomt van een lichaam dat niet alleen sterk is, maar ook goed getoned,